What is the optimal way to
detect pathogenic gene fusions?
detect pathogenic gene fusions?
Next-generation sequencing (NGS) can detect a broad range of genomic alterations1
NGS is a high-throughput genomic sequencing technology that allows for the simultaneous analysis of numerous alterations; NGS can be DNA-based, RNA-based, or both.2,3
Unlike NGS, conventional testing methods, such as RT-PCR, FISH, and IHC, were developed to detect single molecular targets and may fail to uncover the full breadth of genomic alterations.2,4,5
NGS process6
Nucleic acid extraction and isolation
Fragmentation and library creation
Amplifying and sequencing the libraries
DNA-based NGS alone can miss pathogenic gene fusions7
Comprehensive testing with RNA-based NGS, including DNA and RNA sequencing, is recommended to capture what DNA-based NGS alone can miss2,7
DNA-based sequencing can lead to false-negative and false-positive results in a variety of cases, particularly in the detection of gene fusions.7,9
aBased on NGS testing of tissue samples.8
Gain insights into NGS testing platforms

Comprehensive NGS Testing
Gain insights into NGS testing platforms and commercial labs that offer RNA-based NGS testing, including DNA and RNA sequencing.
DOWNLOAD NOW
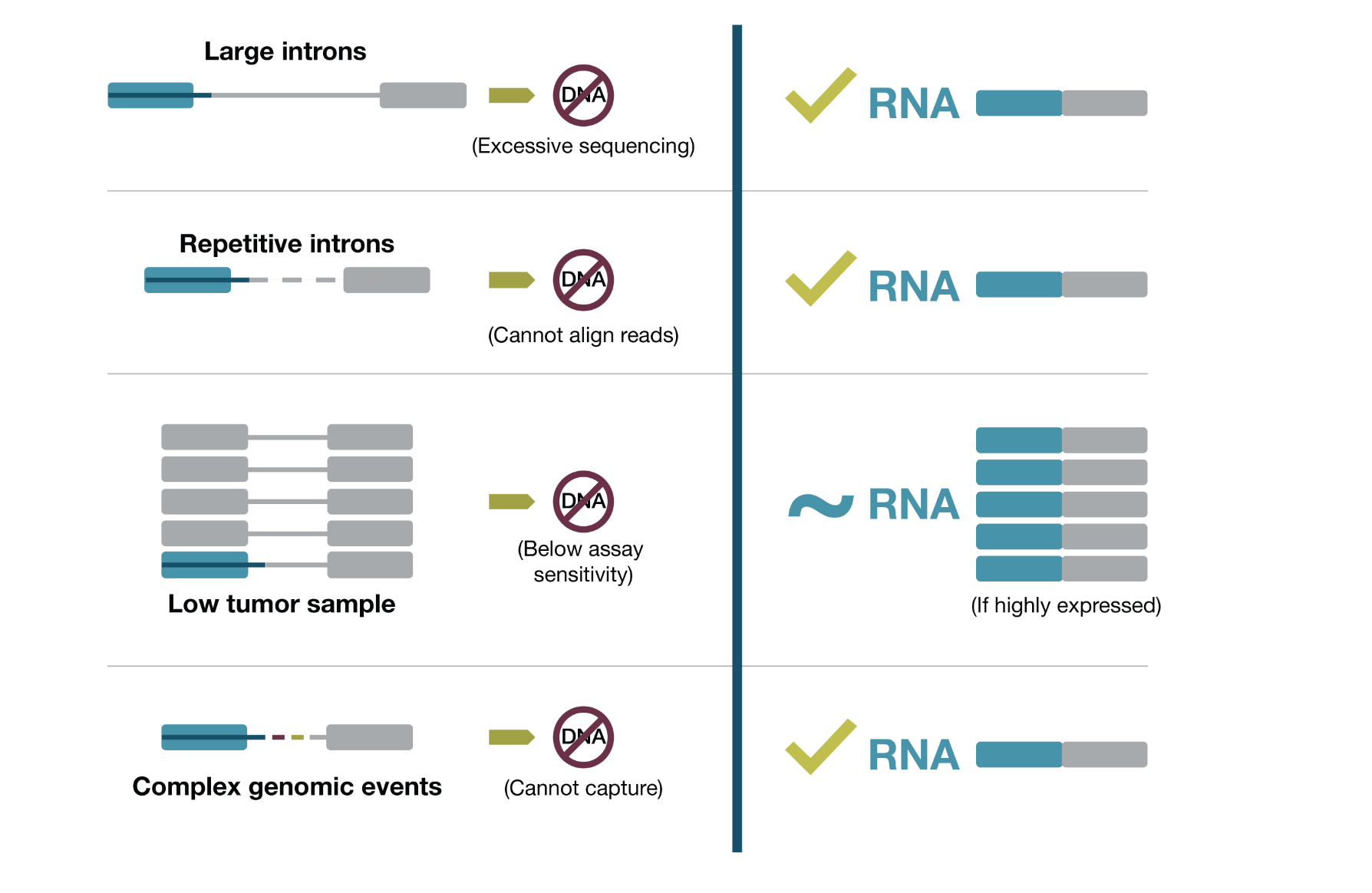
Why is RNA-based NGS, including DNA and RNA sequencing, more comprehensive than DNA-based NGS alone for detecting pathogenic gene fusions?7,10
- RNA-based NGS detects gene expression and many structural variants11
- RNA-based NGS reduces many of the technical challenges involved in sequencing long introns11
- RNA-based NGS enables oncologists to match therapy to the driving fusion, which wouldn't have otherwise been identified, potentially leading to improved clinical responses10,12,13
DNA-based vs RNA-based NGS for fusions14

Watch Dr Chahine discuss
the limitations of DNA NGS
Detecting NRG1 fusions15
- NRG1 fusions are more likely to be missed unless testing with RNA-based NGS
- The diversity of NRG1 fusion partners, breakpoints, and the large intronic regions can make detection more challenging

In a retrospective study by the Memorial Sloan Kettering Cancer Center,
RNA-based NGS detected more NRG1 fusions than DNA-based NGS10
Both DNA-based and RNA-based NGS were performed on 30 IMA samples. Of these, 28 were detected by RNA-based NGS but not DNA-based NGS. The remaining 2 were detected by both. Four additional samples that did not undergo DNA-based NGS were detected by RNA-based NGS.
Detection of NRG1 fusions across tumor types
(MSKCC Experience)10
(MSKCC Experience)10
TEST YOUR KNOWLEDGE
In which cancers are NRG1 fusions most commonly found?

Colorectal cancers

Lung and pancreatic cancers

Hematologic malignancies
NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) include RNA-based NGS16,17
- Pancreatic adenocarcinoma: Tumor/somatic molecular profiling, preferably using NGS assay, is recommended for patients with locally advanced/metastatic disease who are candidates for anti-cancer therapy to identify clinically actionable and/or emerging alterations. These alterations include, but are not limited to, NRG1 fusions. RNA sequencing assays are preferred for detecting RNA fusions because gene fusions are better detected by RNA-based NGS16
- NSCLC: RNA-based NGS may increase novel fusion detection and can be performed concurrently or sequentially with DNA-based NGS. If no identifiable driver oncogenes with DNA-based broad molecular profiling are identified, RNA-based testing is recommended17
ASCO guidelines considered RNA-based methods generally superior for fusion detection across multiple targets.18
NCCN makes no warranties of any kind whatsoever regarding their content, use, or application and disclaims any responsibility for their application or use in any way.

RNA-based NGS on tumor tissue provides the most comprehensive view of the genomic landscape and is a critical step toward optimizing care and potentially improving patient outcomes.16-19
ASCO, American Society of Clinical Oncology; FISH, fluorescence in situ hybridization; IHC, immunohistochemistry; MSKCC, Memorial Sloan Kettering Cancer Center; NCCN, National Comprehensive Cancer Network; NGS, next-generation sequencing; NOS, not otherwise specified; NRG1, neuregulin 1; NSCLC, non–small cell lung cancer; RT-PCR, reverse transcription-polymerase chain reaction.
References: 1. Singh RR. Next-generation sequencing in high-sensitive detection of mutations in tumors: challenges, advances, and applications. J Mol Diagn. 2020;22(8):994-1007. doi:10.1016/j.jmoldx.2020.04.213 2. Bruno R, Fontanini G. Next generation sequencing for gene fusion analysis in lung cancer: a literature review. Diagnostics (Basel). 2020;10(8):521. doi:10.3390/diagnostics10080521 3. Goswami RS, Luthra R, Singh RR, et al. Identification of factors affecting the success of next-generation sequencing testing in solid tumors. Am J Clin Pathol. 2016;145(2):222-237. doi:10.1093/ajcp/aqv023 4. Su D, Zhang D, Chen K, et al. High performance of targeted next generation sequencing on variance detection in clinical tumor specimens in comparison with current conventional methods. J Exp Clin Cancer Res. 2017;36(1):121. doi:10.1186/s13046-017-0591-4 5. Next-generation sequencing testing in oncology. Personalized Medicine in Oncology. Accessed November 18, 2025. https://www.personalizedmedonc.com/supplements/faculty-perspectives-next-generation-sequencing-testing-in-oncology-part-4-of-a-4-part-series/next-generation-sequencing-testing-in-oncology 6. ABM Inc. Next generation sequencing (NGS) - data analysis. Accessed November 18, 2025. https://info.abmgood.com/next-generation-sequencing-ngs-data-analysis 7. Benayed R, Offin M, Mullaney K, et al. High yield of RNA sequencing for targetable kinase fusions in lung adenocarcinomas with no mitogenic driver alteration detected by DNA sequencing and low tumor mutation burden. Clin Cancer Res. 2019;25(15):4712-4722. doi:10.1158/1078-0432.CCR-19-0225 8. Michuda J, Park BH, Cummings AL, et al. Use of clinical RNA-sequencing in the detection of actionable fusions compared to DNA-sequencing alone. J Clin Oncol. 2022;40(16):3077. doi:10.1200/JCO.2022.40.16 9. Heydt C, Wӧlwer CB, Velazquez Camacho O, et al. Detection of gene fusions using targeted next-generation sequencing: a comparative evaluation. BMC Med Genomics. 2021;14(1):62. doi:10.1186/s12920-021-00909-y 10. Benayed R, Liu SV. Neuregulin-1 (NRG1): an emerging tumor-agnostic target. Clinical Care Options: Oncology. Accessed March 2, 2023. https://apps.clinicaloptions.com/oncology/programs/2021/nrg1-fusions/text-module/nrg1-text-module/page-1 11. Mahmoud M, Gobet N, Cruz-Dávalos DI, Mounier N, Dessimoz C, Sedlazeck FJ. Structural variant calling: the long and the short of it. Genome Biol. 2019;20(1):246. doi:10.1186/s13059-019-1828-7 12. Cobain EF, Wu Y-M, Vats P, et al. Assessment of clinical benefit of integrative genomic profiling in advanced solid tumors. JAMA Oncol. 2021;7(4):525-533. doi:10.1001/jamaoncol.2020.7987 13. Hindi I, Shen G, Tan Q, et al. Feasibility and clinical utility of a pan-solid tumor targeted RNA fusion panel: a single center experience. Exp Mol Pathol. 2020;114:104403. doi:10.1016/j.yexmp.2020.104403 14. Davies KD, Aisner DL. Wake up and smell the fusions: single-modality molecular testing misses drivers. Clin Cancer Res. 2019;25(15):4586-4588. doi:10.1158/1078-0432.CCR-19-1361 15. Drilon A, Duruisseaux M, Han J-Y, et al. Clinicopathologic features and response to therapy of NRG1 fusion–driven lung cancers: the eNRGy1 Global Multicenter Registry. J Clin Oncol. 2021;39(25):2791-2802. doi:10.1200/JCO.20.03307 16. Referenced with permission from the NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) for Pancreatic Adenocarcinoma V.2.2025. © National Comprehensive Cancer Network, Inc. 2025. All rights reserved. Accessed August 27, 2025. To view the most recent and complete version of the guideline, go online to NCCN.org. NCCN makes no warranties of any kind whatsoever regarding their content, use or application and disclaims any responsibility for their application or use in any way. 17. Referenced with permission from the NCCN Clinical Practice Guidelines in Oncology (NCCN Guidelines®) for Non-Small Cell Lung Cancer V.3.2026. © National Comprehensive Cancer Network, Inc. 2025. All rights reserved. Accessed January 15, 2026. To view the most recent and complete version of the guideline, go online to NCCN.org. NCCN makes no warranties of any kind whatsoever regarding their content, use or application and disclaims any responsibility for their application or use in any way. 18. Chakravarty D, Johnson Amber, Sklar J, et al. Somatic genomic testing in patients with metastatic or advanced cancer: ASCO provisional clinical opinion. J Clin Oncol. 2022; 40:1231-1258. doi: 10.1200/JCO.21.02767 19. Iams WT, Mackay M, Ben-Shachar R, et al. Concurrent tissue and circulating tumor DNA molecular profiling to detect guideline-based targeted mutations in a multicenter cohort. JAMA Netw Open. 2024;7(1):e2351700. doi:10.1001/jamanetworkopen.2023.51700


